annnet
annnet (Annotated Network) is a unified, high‑expressivity graph platform that brings anndata‑style, annotated containers to networks, multilayer structures, and hypergraphs. It targets systems biology, network biology, omics integration, computational social science, and any domain needing fully flexible graph semantics with modern, stable storage and interoperability.
Why annnet
Many real-world networks are structurally heterogeneous and context-dependent. A single dataset may combine directed and undirected interactions, signed edges, higher-order relations, and multiple experimental or temporal conditions. Standard graph libraries handle topology and algorithms well, but typically treat attributes as flat, per-object dictionaries without schema, indexing, or efficient bulk operations. This makes it difficult to manage annotations, compare conditions, or preserve structure across analysis steps.
annnet addresses this by defining a single container that keeps graph topology, annotation tables, and graph views aligned. The goal is not to replace existing graph libraries, but to provide a consistent data model that can express complex networks and still interoperate with established tooling.
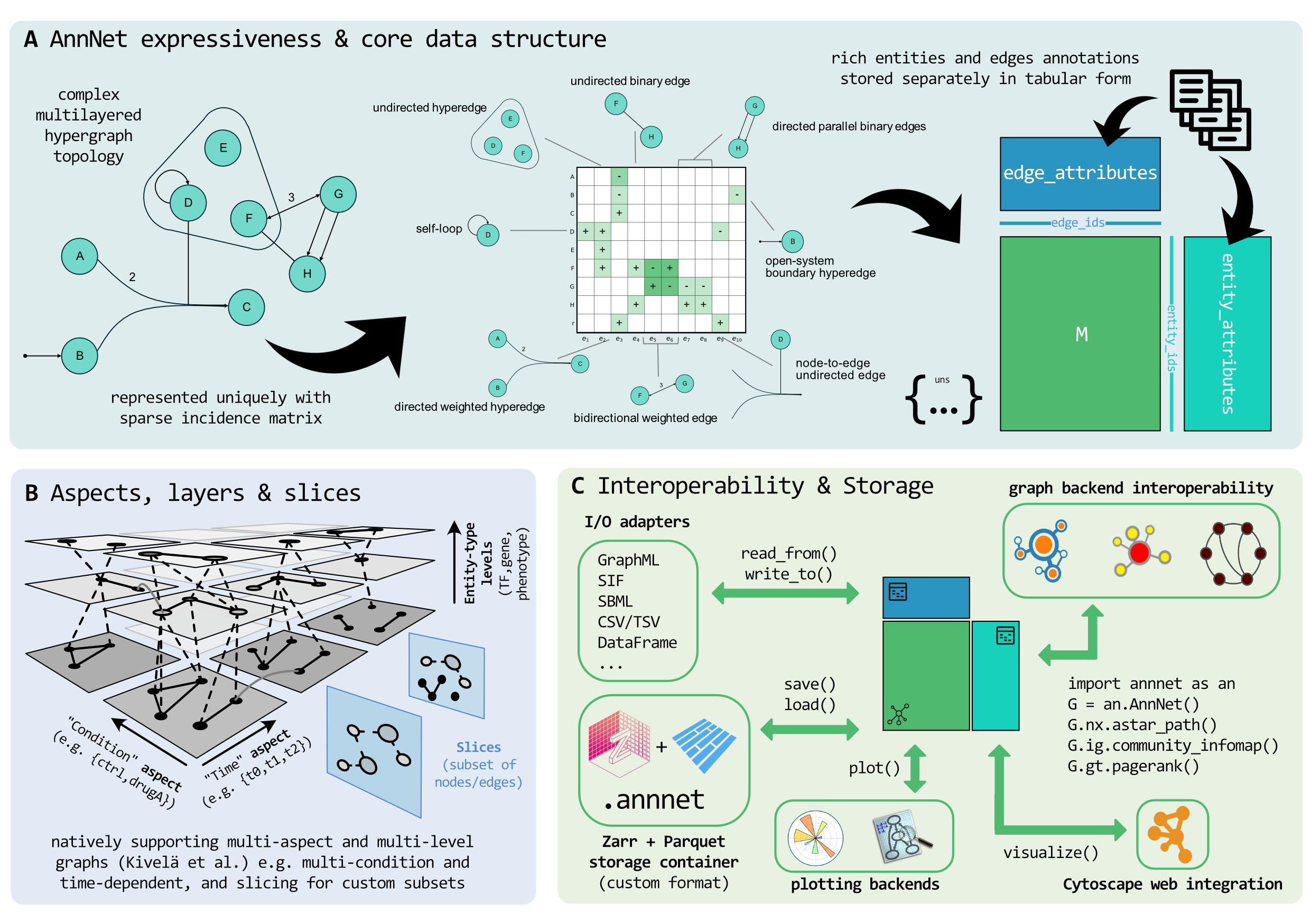
At the core is a sparse incidence-based representation that supports mixed graph types within the same object. Edges are first-class entities with stable identifiers, and graph type (directed, undirected, hyperedge) is a property of each edge rather than the container. Around this core, annnet organizes metadata as typed tables and exposes higher-level constructs such as slices and multilayer structure without duplicating the underlying graph.
-
One object for heterogeneous graphs
Represent simple graphs, digraphs, signed edges, hyperedges, self-loops, parallel edges, and edge-entity relations without switching data models.
-
Typed annotation tables
Store vertex, edge, slice, layer, and edge-slice metadata in indexed tabular structures instead of flat per-object dictionaries.
-
Named slices for conditions and views
Define condition-specific or context-specific graph views without copying the full topology, with optional per-slice edge-weight overrides.
-
Multilayer network support
Work with aspects, layer tuples, vertex-layer membership, intra-layer edges, inter-layer edges, and coupling structure as part of the core model.
-
Interoperability without losing the source of truth
Convert lazily to existing graph libraries when needed, while keeping annnet as the canonical representation.
-
Native, loss-aware storage
Persist topology, annotations, and metadata together in a format designed for round-trip fidelity and scalable IO.
What annnet is built for
annnet is most useful in settings where graph structure is only one part of the data model, and where annotations, conditions, or multiple representations must be handled explicitly. The design follows patterns that have proven useful in other domains (for example, matrix-plus-annotation containers in omics, in AnnData), but adapts them to graphs with heterogeneous topology.
Instead of encoding these requirements through ad hoc conventions or multiple loosely coupled objects, annnet keeps them within a single, consistent representation that can still be exported or adapted when needed.
-
Systems biology and omics integration
Model regulatory, signaling, metabolic, and cross-modal networks with typed metadata and multiple experimental contexts.
-
Condition-specific and temporal networks
Keep one shared graph with named slices for perturbations, time points, cohorts, or filtered analytical views.
-
Multimodal and layered data
Represent networks across modalities, resolutions, or time as explicit multilayer objects rather than ad hoc conventions.
-
Exchange with existing tooling
Move between annnet and standard graph ecosystems, file formats, and ML pipelines without flattening the original structure too early.
Documentation
The documentation is structured around how the package is typically used: first understanding the model, then working through examples, and finally consulting the API in detail. Check out the documentation sections below:
-
Tutorials and notebooks
End-to-end examples showing graph construction, annotation workflows, slices, multilayer models, and interoperability.
-
Concepts and design
Explanations of the incidence-matrix core, annotation system, slices, multilayer formalism, adapters, and storage model.
-
API reference
Detailed reference for the object model, bulk APIs, IO, utilities, and public entry points.
-
Community
Contribution guidance for documentation, package development, and project standards.
Start with the workflow you need
The entry points below correspond to common usage patterns. Installation is the minimal setup path, the quickstart introduces the core object and data model, and the tutorial walks through a complete example with annotations, hypergraph structure, and layers.
Install annnet, build a toy AnnNet graph, or inspect a full biological example
Start with installation if you are setting up the package, the quickstart if you want the core object model, or the tutorial if you want a complete worked example with annotations and graph structure.