Pathway activity inference in bulk RNA-seq
Pau Badia-i-Mompel
Heidelberg UniversiySource:
vignettes/pw_bk.Rmd
pw_bk.RmdBulk RNA-seq yield many molecular readouts that are hard to interpret by themselves. One way of summarizing this information is by inferring pathway activities from prior knowledge.
In this notebook we showcase how to use decoupleR for
pathway activity inference with a bulk RNA-seq data-set where the
transcription factor FOXA2 was knocked out in pancreatic cancer cell
lines.
The data consists of 3 Wild Type (WT) samples and 3 Knock Outs (KO). They are freely available in GEO.
Loading the data-set
Here we used an already processed bulk RNA-seq data-set. We provide
the normalized log-transformed counts, the experimental design meta-data
and the Differential Expressed Genes (DEGs) obtained using
limma.
For this example we use limma but we could have used
DeSeq2, edgeR or any other statistical
framework. decoupleR requires a gene level statistic to perform
enrichment analysis but it is agnostic of how it was generated. However,
we do recommend to use statistics that include the direction of change
and its significance, for example the t-value obtained for
limma(t) or
DeSeq2(stat). edgeR does not return such
statistic but we can create our own by weighting the obtained logFC by
pvalue with this formula: -log10(pvalue) * logFC.
We can open the data like this:
inputs_dir <- system.file("extdata", package = "decoupleR")
data <- readRDS(file.path(inputs_dir, "bk_data.rds"))From data we can extract the mentioned information. Here
we see the normalized log-transformed counts:
# Remove NAs and set row names
counts <- data$counts %>%
dplyr::mutate_if(~ any(is.na(.x)),
~ dplyr::if_else(is.na(.x), 0, .x)) %>%
tibble::column_to_rownames(var = "gene") %>%
as.matrix()
head(counts)
#> PANC1.WT.Rep1 PANC1.WT.Rep2 PANC1.WT.Rep3 PANC1.FOXA2KO.Rep1
#> NOC2L 10.052588 11.949123 12.057774 12.312291
#> PLEKHN1 7.535115 8.125993 8.714880 8.048196
#> PERM1 6.281242 6.424582 6.589668 6.293285
#> ISG15 10.938252 11.469081 11.425415 11.549986
#> AGRN 6.956335 7.196108 7.522550 7.061549
#> C1orf159 9.546224 9.788721 9.794589 9.850830
#> PANC1.FOXA2KO.Rep2 PANC1.FOXA2KO.Rep3
#> NOC2L 12.139918 11.494205
#> PLEKHN1 8.290154 8.621239
#> PERM1 6.486136 6.775344
#> ISG15 11.371464 11.178157
#> AGRN 7.485534 7.071555
#> C1orf159 9.988069 9.965357The design meta-data:
design <- data$design
design
#> # A tibble: 6 × 2
#> sample condition
#> <chr> <chr>
#> 1 PANC1.WT.Rep1 PANC1.WT
#> 2 PANC1.WT.Rep2 PANC1.WT
#> 3 PANC1.WT.Rep3 PANC1.WT
#> 4 PANC1.FOXA2KO.Rep1 PANC1.FOXA2KO
#> 5 PANC1.FOXA2KO.Rep2 PANC1.FOXA2KO
#> 6 PANC1.FOXA2KO.Rep3 PANC1.FOXA2KOAnd the results of limma, of which we are interested in
extracting the obtained t-value from the contrast:
PROGENy model
PROGENy is a comprehensive resource containing a curated collection of pathways and their target genes, with weights for each interaction. For this example we will use the human weights (other organisms are available) and we will use the top 500 responsive genes ranked by p-value. Here is a brief description of each pathway:
- Androgen: involved in the growth and development of the male reproductive organs.
- EGFR: regulates growth, survival, migration, apoptosis, proliferation, and differentiation in mammalian cells
- Estrogen: promotes the growth and development of the female reproductive organs.
- Hypoxia: promotes angiogenesis and metabolic reprogramming when O2 levels are low.
- JAK-STAT: involved in immunity, cell division, cell death, and tumor formation.
- MAPK: integrates external signals and promotes cell growth and proliferation.
- NFkB: regulates immune response, cytokine production and cell survival.
- p53: regulates cell cycle, apoptosis, DNA repair and tumor suppression.
- PI3K: promotes growth and proliferation.
- TGFb: involved in development, homeostasis, and repair of most tissues.
- TNFa: mediates haematopoiesis, immune surveillance, tumour regression and protection from infection.
- Trail: induces apoptosis.
- VEGF: mediates angiogenesis, vascular permeability, and cell migration.
- WNT: regulates organ morphogenesis during development and tissue repair.
To access it we can use decoupleR:
net <- decoupleR::get_progeny(organism = 'human',
top = 500)
net
#> # A tibble: 7,000 × 4
#> source target weight p_value
#> <chr> <chr> <dbl> <dbl>
#> 1 Androgen TMPRSS2 11.5 2.38e-47
#> 2 Androgen NKX3-1 10.6 2.21e-44
#> 3 Androgen MBOAT2 10.5 4.63e-44
#> 4 Androgen KLK2 10.2 1.94e-40
#> 5 Androgen SARG 11.4 2.79e-40
#> 6 Androgen SLC38A4 7.36 1.25e-39
#> 7 Androgen MTMR9 6.13 2.53e-38
#> 8 Androgen ZBTB16 10.6 1.57e-36
#> 9 Androgen KCNN2 9.47 7.71e-36
#> 10 Androgen OPRK1 -5.63 1.11e-35
#> # ℹ 6,990 more rowsActivity inference with Multivariate Linear Model (MLM)
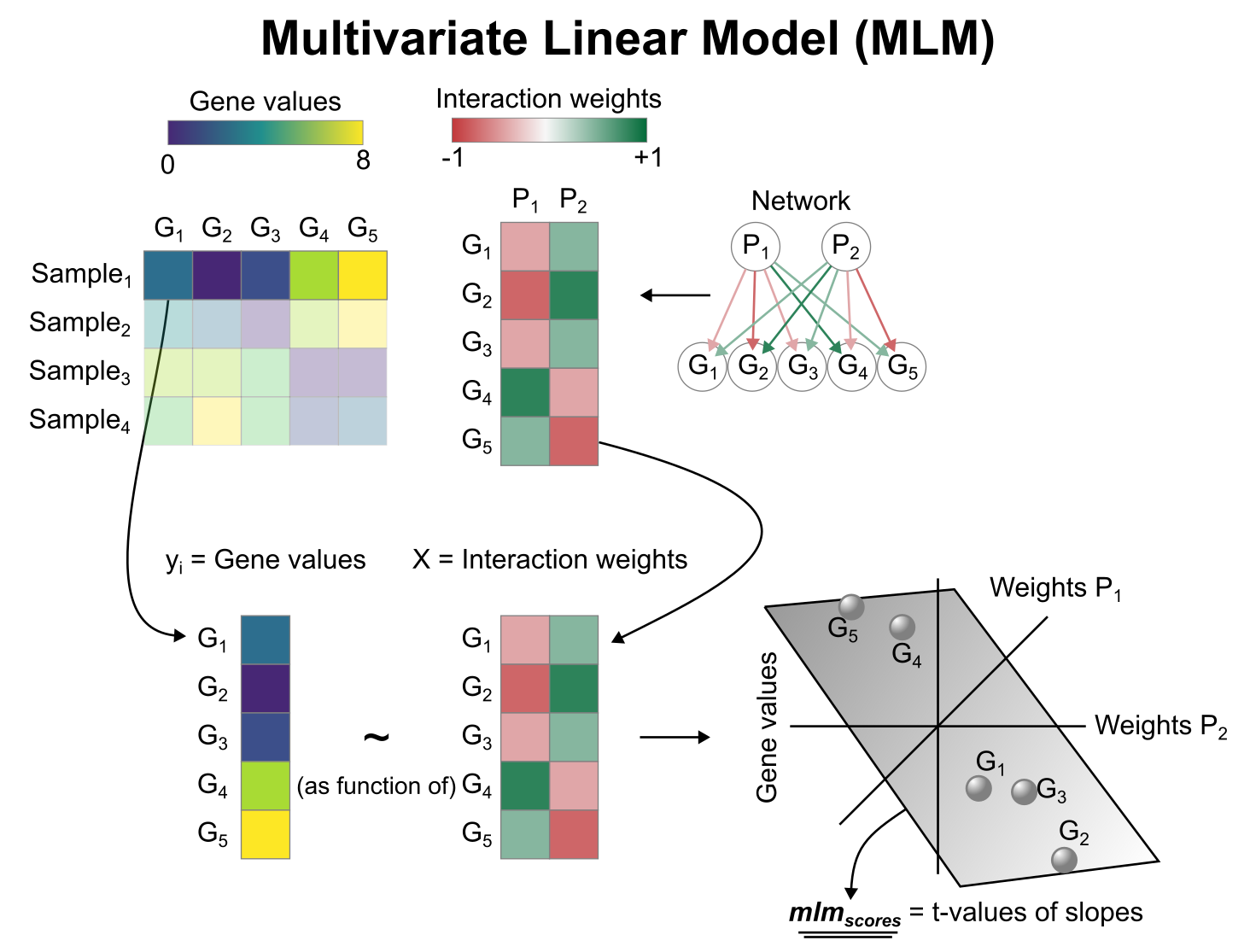
To infer pathway enrichment scores we will run the Multivariate
Linear Model (mlm) method. For each sample in our dataset
(mat), it fits a linear model that predicts the observed
gene expression based on all pathways’ Pathway-Gene interactions
weights. Once fitted, the obtained t-values of the slopes are the
scores. If it is positive, we interpret that the pathway is active and
if it is negative we interpret that it is inactive.
To run decoupleR methods, we need an input matrix
(mat), an input prior knowledge network/resource
(net), and the name of the columns of net that we want to
use.
# Run mlm
sample_acts <- decoupleR::run_mlm(mat = counts,
net = net,
.source = 'source',
.target = 'target',
.mor = 'weight',
minsize = 5)
sample_acts
#> # A tibble: 84 × 5
#> statistic source condition score p_value
#> <chr> <chr> <chr> <dbl> <dbl>
#> 1 mlm Androgen PANC1.WT.Rep1 -0.692 0.489
#> 2 mlm EGFR PANC1.WT.Rep1 -0.0414 0.967
#> 3 mlm Estrogen PANC1.WT.Rep1 -0.361 0.718
#> 4 mlm Hypoxia PANC1.WT.Rep1 -2.06 0.0393
#> 5 mlm JAK-STAT PANC1.WT.Rep1 -0.166 0.868
#> 6 mlm MAPK PANC1.WT.Rep1 -0.509 0.611
#> 7 mlm NFkB PANC1.WT.Rep1 -2.84 0.00447
#> 8 mlm PI3K PANC1.WT.Rep1 3.53 0.000423
#> 9 mlm TGFb PANC1.WT.Rep1 -1.43 0.152
#> 10 mlm TNFa PANC1.WT.Rep1 2.06 0.0395
#> # ℹ 74 more rowsVisualization
From the obtained results we will observe the obtained activities per sample in a heat-map:
# Transform to wide matrix
sample_acts_mat <- sample_acts %>%
tidyr::pivot_wider(id_cols = 'condition',
names_from = 'source',
values_from = 'score') %>%
tibble::column_to_rownames('condition') %>%
as.matrix()
# Scale per feature
sample_acts_mat <- scale(sample_acts_mat)
# Color scale
colors <- rev(RColorBrewer::brewer.pal(n = 11, name = "RdBu"))
colors.use <- grDevices::colorRampPalette(colors = colors)(100)
my_breaks <- c(seq(-2, 0, length.out = ceiling(100 / 2) + 1),
seq(0.05,2, length.out = floor(100 / 2)))
# Plot
pheatmap::pheatmap(mat = sample_acts_mat,
color = colors.use,
border_color = "white",
breaks = my_breaks,
cellwidth = 20,
cellheight = 20,
treeheight_row = 20,
treeheight_col = 20)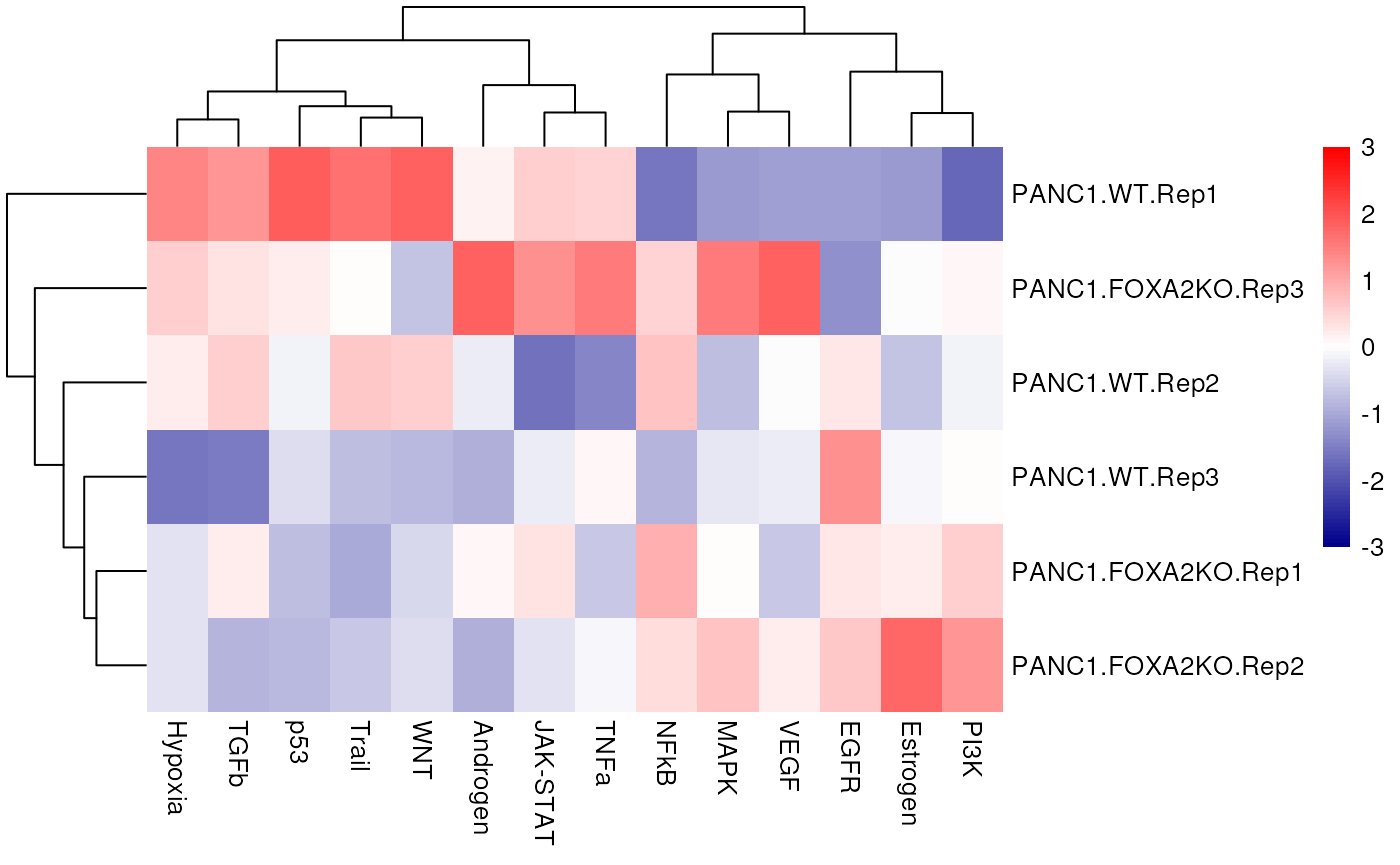
We can also infer pathway activities from the t-values of the DEGs between KO and WT:
# Run mlm
contrast_acts <- decoupleR::run_mlm(mat =deg,
net = net,
.source = 'source',
.target = 'target',
.mor = 'weight',
minsize = 5)
contrast_acts
#> # A tibble: 14 × 5
#> statistic source condition score p_value
#> <chr> <chr> <chr> <dbl> <dbl>
#> 1 mlm Androgen t -0.219 8.27e- 1
#> 2 mlm EGFR t -0.439 6.61e- 1
#> 3 mlm Estrogen t 3.95 7.70e- 5
#> 4 mlm Hypoxia t 0.171 8.64e- 1
#> 5 mlm JAK-STAT t 5.91 3.47e- 9
#> 6 mlm MAPK t 13.0 4.32e-38
#> 7 mlm NFkB t 1.36 1.75e- 1
#> 8 mlm PI3K t 5.65 1.65e- 8
#> 9 mlm TGFb t -0.656 5.12e- 1
#> 10 mlm TNFa t 1.93 5.36e- 2
#> 11 mlm Trail t -2.03 4.26e- 2
#> 12 mlm VEGF t 2.82 4.80e- 3
#> 13 mlm WNT t -1.49 1.35e- 1
#> 14 mlm p53 t -4.80 1.60e- 6Let’s show the changes in activity between KO and WT:
# Plot
colors <- rev(RColorBrewer::brewer.pal(n = 11, name = "RdBu")[c(2, 10)])
p <- ggplot2::ggplot(data = contrast_acts,
mapping = ggplot2::aes(x = stats::reorder(source, score),
y = score)) +
ggplot2::geom_bar(mapping = ggplot2::aes(fill = score),
color = "black",
stat = "identity") +
ggplot2::scale_fill_gradient2(low = colors[1],
mid = "whitesmoke",
high = colors[2],
midpoint = 0) +
ggplot2::theme_minimal() +
ggplot2::theme(axis.title = element_text(face = "bold", size = 12),
axis.text.x = ggplot2::element_text(angle = 45,
hjust = 1,
size = 10,
face = "bold"),
axis.text.y = ggplot2::element_text(size = 10,
face = "bold"),
panel.grid.major = element_blank(),
panel.grid.minor = element_blank()) +
ggplot2::xlab("Pathways")
p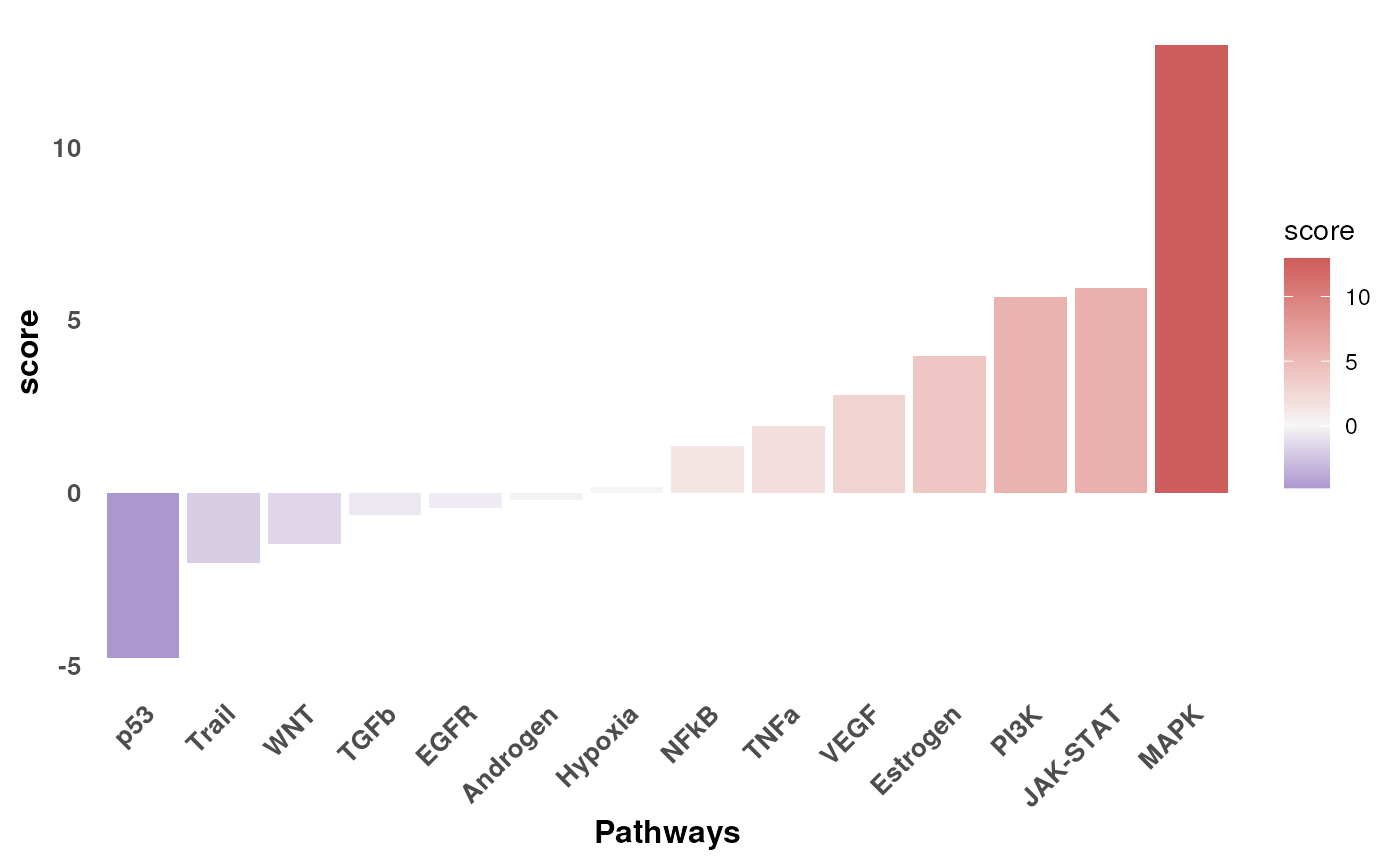
The pathway p53 and Trail are deactivated in KO when compared to WT, while MAPKK and JAK-STAT and seem to be activated.
We can further visualize the most responsive genes in each pathway along their t-values to interpret the results. For example, let’s see the genes that are belong to the MAPK pathway:
pathway <- 'MAPK'
df <- net %>%
dplyr::filter(source == pathway) %>%
dplyr::arrange(target) %>%
dplyr::mutate(ID = target,
color = "3") %>%
tibble::column_to_rownames('target')
inter <- sort(dplyr::intersect(rownames(deg), rownames(df)))
df <- df[inter, ]
df['t_value'] <- deg[inter, ]
df <- df %>%
dplyr::mutate(color = dplyr::if_else(weight > 0 & t_value > 0, '1', color)) %>%
dplyr::mutate(color = dplyr::if_else(weight > 0 & t_value < 0, '2', color)) %>%
dplyr::mutate(color = dplyr::if_else(weight < 0 & t_value > 0, '2', color)) %>%
dplyr::mutate(color = dplyr::if_else(weight < 0 & t_value < 0, '1', color))
colors <- rev(RColorBrewer::brewer.pal(n = 11, name = "RdBu")[c(2, 10)])
p <- ggplot2::ggplot(data = df,
mapping = ggplot2::aes(x = weight,
y = t_value,
color = color)) +
ggplot2::geom_point(size = 2.5,
color = "black") +
ggplot2::geom_point(size = 1.5) +
ggplot2::scale_colour_manual(values = c(colors[2], colors[1], "grey")) +
ggrepel::geom_label_repel(mapping = ggplot2::aes(label = ID)) +
ggplot2::theme_minimal() +
ggplot2::theme(legend.position = "none") +
ggplot2::geom_vline(xintercept = 0, linetype = 'dotted') +
ggplot2::geom_hline(yintercept = 0, linetype = 'dotted') +
ggplot2::ggtitle(pathway)
p
#> Warning: ggrepel: 447 unlabeled data points (too many overlaps). Consider
#> increasing max.overlaps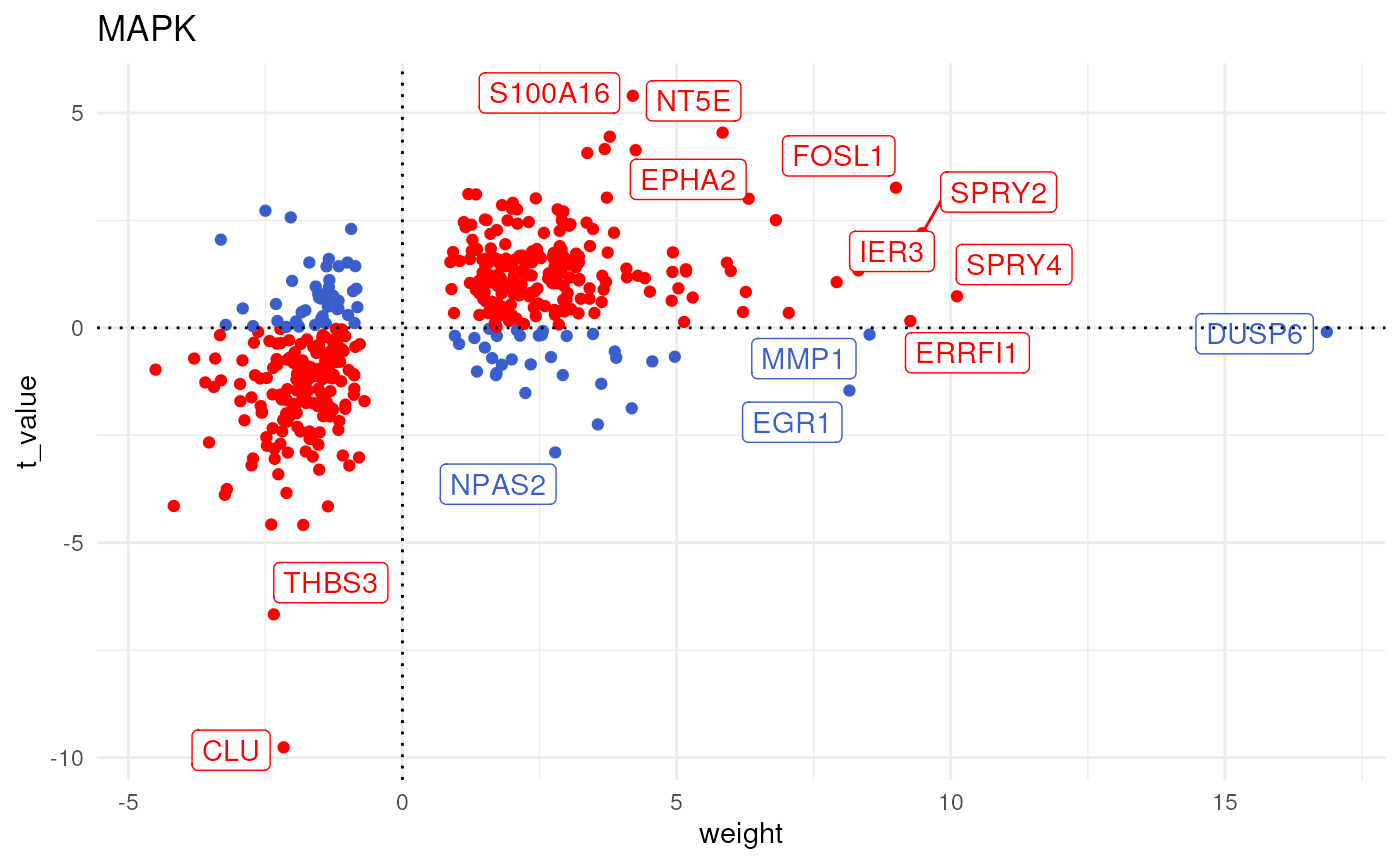
The pathway seems to be active since the majority of target genes with positive weights have positive t-values (1st quadrant), and the majority of the ones with negative weights have negative t-values (3d quadrant).
Session information
#> ─ Session info ───────────────────────────────────────────────────────────────────────────────────────────────────────
#> setting value
#> version R version 4.4.1 (2024-06-14)
#> os Ubuntu 22.04.5 LTS
#> system x86_64, linux-gnu
#> ui X11
#> language en
#> collate en_US.UTF-8
#> ctype en_US.UTF-8
#> tz UTC
#> date 2024-10-19
#> pandoc 3.4 @ /usr/bin/ (via rmarkdown)
#>
#> ─ Packages ───────────────────────────────────────────────────────────────────────────────────────────────────────────
#> package * version date (UTC) lib source
#> backports 1.5.0 2024-05-23 [1] RSPM
#> BiocManager 1.30.25 2024-08-28 [1] RSPM
#> BiocParallel 1.38.0 2024-04-30 [1] Bioconduc~
#> BiocStyle * 2.32.1 2024-06-16 [1] Bioconduc~
#> bit 4.5.0 2024-09-20 [1] RSPM
#> bit64 4.5.2 2024-09-22 [1] RSPM
#> bookdown 0.41 2024-10-16 [1] RSPM
#> bslib 0.8.0 2024-07-29 [1] RSPM
#> cachem 1.1.0 2024-05-16 [1] RSPM
#> cellranger 1.1.0 2016-07-27 [1] RSPM
#> checkmate 2.3.2 2024-07-29 [1] RSPM
#> cli 3.6.3 2024-06-21 [1] RSPM
#> codetools 0.2-20 2024-03-31 [2] CRAN (R 4.4.1)
#> colorspace 2.1-1 2024-07-26 [1] RSPM
#> crayon 1.5.3 2024-06-20 [1] RSPM
#> curl 5.2.3 2024-09-20 [1] RSPM
#> decoupleR * 2.9.7 2024-10-19 [1] Bioconductor
#> desc 1.4.3 2023-12-10 [1] RSPM
#> digest 0.6.37 2024-08-19 [1] RSPM
#> dplyr * 1.1.4 2023-11-17 [1] RSPM
#> evaluate 1.0.1 2024-10-10 [1] RSPM
#> fansi 1.0.6 2023-12-08 [1] RSPM
#> farver 2.1.2 2024-05-13 [1] RSPM
#> fastmap 1.2.0 2024-05-15 [1] RSPM
#> fs 1.6.4 2024-04-25 [1] RSPM
#> generics 0.1.3 2022-07-05 [1] RSPM
#> ggplot2 * 3.5.1 2024-04-23 [1] RSPM
#> ggrepel * 0.9.6 2024-09-07 [1] RSPM
#> glue 1.8.0 2024-09-30 [1] RSPM
#> gtable 0.3.5 2024-04-22 [1] RSPM
#> highr 0.11 2024-05-26 [1] RSPM
#> hms 1.1.3 2023-03-21 [1] RSPM
#> htmltools 0.5.8.1 2024-04-04 [1] RSPM
#> htmlwidgets 1.6.4 2023-12-06 [1] RSPM
#> httr 1.4.7 2023-08-15 [1] RSPM
#> igraph 2.0.3 2024-03-13 [1] RSPM
#> jquerylib 0.1.4 2021-04-26 [1] RSPM
#> jsonlite 1.8.9 2024-09-20 [1] RSPM
#> knitr 1.48 2024-07-07 [1] RSPM
#> labeling 0.4.3 2023-08-29 [1] RSPM
#> later 1.3.2 2023-12-06 [1] RSPM
#> lattice 0.22-6 2024-03-20 [2] CRAN (R 4.4.1)
#> lifecycle 1.0.4 2023-11-07 [1] RSPM
#> logger 0.3.0 2024-03-05 [1] RSPM
#> lubridate 1.9.3 2023-09-27 [1] RSPM
#> magrittr 2.0.3 2022-03-30 [1] RSPM
#> Matrix 1.7-1 2024-10-18 [1] RSPM (R 4.4.0)
#> munsell 0.5.1 2024-04-01 [1] RSPM
#> OmnipathR 3.12.4 2024-10-02 [1] Bioconduc~
#> parallelly 1.38.0 2024-07-27 [1] RSPM
#> pheatmap * 1.0.12 2019-01-04 [1] RSPM
#> pillar 1.9.0 2023-03-22 [1] RSPM
#> pkgconfig 2.0.3 2019-09-22 [1] RSPM
#> pkgdown 2.1.1 2024-09-17 [1] RSPM
#> prettyunits 1.2.0 2023-09-24 [1] RSPM
#> progress 1.2.3 2023-12-06 [1] RSPM
#> purrr 1.0.2 2023-08-10 [1] RSPM
#> R6 2.5.1 2021-08-19 [1] RSPM
#> ragg 1.3.3 2024-09-11 [1] RSPM
#> rappdirs 0.3.3 2021-01-31 [1] RSPM
#> RColorBrewer 1.1-3 2022-04-03 [1] RSPM
#> Rcpp 1.0.13 2024-07-17 [1] RSPM
#> readr 2.1.5 2024-01-10 [1] RSPM
#> readxl 1.4.3 2023-07-06 [1] RSPM
#> rlang 1.1.4 2024-06-04 [1] RSPM
#> rmarkdown 2.28 2024-08-17 [1] RSPM
#> rvest 1.0.4 2024-02-12 [1] RSPM
#> sass 0.4.9 2024-03-15 [1] RSPM
#> scales 1.3.0 2023-11-28 [1] RSPM
#> selectr 0.4-2 2019-11-20 [1] RSPM
#> sessioninfo 1.2.2 2021-12-06 [1] RSPM
#> stringi 1.8.4 2024-05-06 [1] RSPM
#> stringr 1.5.1 2023-11-14 [1] RSPM
#> systemfonts 1.1.0 2024-05-15 [1] RSPM
#> textshaping 0.4.0 2024-05-24 [1] RSPM
#> tibble * 3.2.1 2023-03-20 [1] RSPM
#> tidyr * 1.3.1 2024-01-24 [1] RSPM
#> tidyselect 1.2.1 2024-03-11 [1] RSPM
#> timechange 0.3.0 2024-01-18 [1] RSPM
#> tzdb 0.4.0 2023-05-12 [1] RSPM
#> utf8 1.2.4 2023-10-22 [1] RSPM
#> vctrs 0.6.5 2023-12-01 [1] RSPM
#> vroom 1.6.5 2023-12-05 [1] RSPM
#> withr 3.0.1 2024-07-31 [1] RSPM
#> xfun 0.48 2024-10-03 [1] RSPM
#> xml2 1.3.6 2023-12-04 [1] RSPM
#> yaml 2.3.10 2024-07-26 [1] RSPM
#>
#> [1] /usr/local/lib/R/site-library
#> [2] /usr/local/lib/R/library
#>
#> ──────────────────────────────────────────────────────────────────────────────────────────────────────────────────────